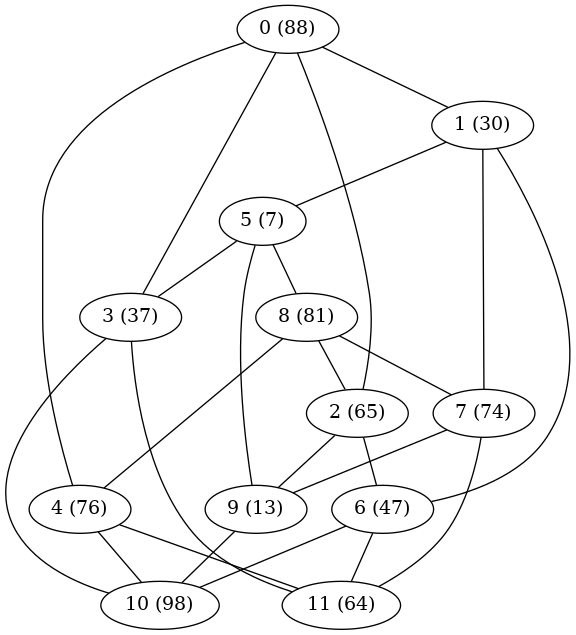
In a previous post, I described a genetic algorithm for a problem posted on OR Stack Exchange. The problem involved grouping users and servers so as to maximize the total quality of service to all users. Every user in a cluster is served by every server in the cluster, but servers in other clusters interfere to some degree with the user's service.
The person who posted that problem recently posted another variation on OR SE. The new version of the problem is nearly, but not quite, identical to the previous version. "Servers" are now termed "transmitters", but the problem remains clustering transmitters and users, with limits on how many transmitters (but not how many users) belong to each cluster, and with the same objective function. The new version may be slightly easier thanks to one new restriction.
The user must belong to a cluster that has the transmitter providing the highest weight for the given user.
Thus the assignment of a transmitter to a cluster immediately implies the assignment to that cluster of all users for whom the specified transmitter has highest weight. (I will assume that ties for highest weight either do not occur or are broken whimsically.)
I put together two different mixed-integer programming (MIP) models for the problem. Both are rather complicated, in part because the objective function needs to be linearized. They are too big to squeeze into a blog post, so I put them in a PDF file. The models differ in the use of binary variables to assign transmitters. In the first model, a binary variable $x_{t,t'}$ is 1 if transmitters $t$ and $t'$ belong to the same cluster. In the second, a binary variable $x_{t,c}$ is 1 if transmitter $t$ belongs to cluster $c$. I coded both models in Java and ran them (using CPLEX as the MIP solver) on a few test problems with dimensions similar to what the original post contained (20 transmitters, 60 users). The first model was consistently smaller in size than the second and produced tighter bounds within whatever time limit I gave CPLEX.
The OR SE question also mentioned heuristics, so I coded up a couple of reasonably obvious ones. The first is a greedy construction heuristic. It initially puts each transmitter in a separate cluster, then repeatedly merges the pair of clusters that either produces the best improvement in the overall objective function. If no further improvement is possible and there are still more clusters than the number allowed, it merges whichever pair does the least damage to the objective. In all cases, mergers occur only if the merged cluster is within the allowed size limit.
The second heuristic is a pairwise swap heuristic. Given the (presumably feasible) solution from the first heuristic, it considers all pairs of transmitters belong to different clusters and, for each pair, evaluates the objective impact of swapping the two transmitters and performs the swap if it is beneficial. Note that swapping preserves the sizes of both clusters, so there is no question whether it is feasible. The heuristic keeps looping through pairs of transmitters until it is unable to find a useful swap.
In my experiments, I used the best solution from the second heuristic to "hot start" the first MIP model. (I ran into some difficulties getting the second MIP model to accept it as a starting solution, and frankly was not motivated enough to bother investigating the issue.) The results from one computational test (20 transmitters, 60 users, maximum cluster size 5) are fairly representative of what I saw in other tests. The construction heuristic got a solution with objective value 26.98 in 0.1 seconds on my desktop PC. The improvement heuristic ran for about 0.25 seconds and improved the objective value to 29.05. Running the first (better) MIP model using CPLEX 20.1 with a 10 minute time limit, starting from that solution, CPLEX improved the objective value to 29.62 with a final upper bound around 31.18. So the heuristic solution, which was obtained very quickly, was within 7% of optimal based on the bound and within 2% of the last solution CPLEX got within the 10 minute time limit.
Two caveats apply here. First, I did a minimal amount of testing. Second, and possibly more important, I randomly generated the transmitter-user weights using a uniform distribution. Having weights that follow a different distribution might affect both heuristic and model performance.
The Java code for both heuristics and both MIP models can be downloaded from my repository. It requires a recent version of Java (14 or later) since it uses the Record class, and it of course requires CPLEX (unless you want to delete that part and just play with the heuristics), but has no other dependencies.